Clinical Case Reports and Clinical Study
OPEN ACCESS | Volume 13 - Issue 1 - 2026
ISSN No: 2766-8614 | Journal DOI: 10.61148/2766-8614/JCCRCS
Sorush Niknamian
MD Ph.D. MPH Dr.PH, Military Medicine, Liberty University (LU), VA, USA
Fellow Member of International Society of Infectious Disease (ISID)
*Corresponding author: Sorush Niknamian, Military Medicine Dep, Liberty University, USA
Received: February 19, 2021
Accepted: March 18, 2021
Published: March 23, 2021
Citation: Niknamian S. “ On the Neglected Shifting balance theory, Bateson–Dobzhansky–Muller model & Quantum evolution plus the Role of Mitochondrial Membrane Potential (MMP) Impact on COVID-19”. Clinical Case Reports and Clinical Study, 3(1) ; DOI: 10.61148/2766-8614/JCCRCS/036
Copyright: © 2021 Sorush Niknamian. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Background: Approximately 80% of all viruses are RNA viruses and they contain their specific RNA helicases. Defective RNA helicases have been linked to infectious diseases (Viral Infections).
Materials and Methods: The articles have gone through many types of research from the beginning of the epidemic of Coronaviruses through history and we introduced the neglected hypothesis of Shifting balance theory, Bateson–Dobzhansky–Muller model & Quantum evolution. In the ancestral population, the genotype is AABB. When two populations become isolated from each other, new mutations can arise. In one population A evolves into a, and in the other B evolves into b. When the two populations hybridize it is the first time A and B interact with each other. When these alleles are incompatible, we speak of Dobzhansky–Muller incompatibilities plus the role of MMA in mitochondria in spreading SARS-CoV-19 through populations and the result of an infection in COVID-19.
Results: In viruses specifically COVID-19, Ribosomal Frameshift is programmed to allows the virus to encode multiple types of proteins from the same mRNA. HIV-1 (human immunodeficiency virus), RSV (Rous sarcoma virus), and all types of influenza viruses use Ribosomal Frameshift. they rely on frameshifting to create a proper ratio of normal translation and trans-frame (encoded by frameshifted sequence) proteins. Notably, its use in viruses is primarily for compacting more genetic information into a shorter amount of genetic material.
Conclusion: to find the genome sequence of COVID-19 we also used Nanopore sequencing that introduced and manufactured by Oxford scientists, due to differences in the action of infection in the host, we could not reach any results since the Novel Virus has not a stable genome (which is quite dynamic) since through our deep research, each virus contains its specific genome sequencing and we cannot claim that COVID-19 has one specific genome sequence like MERS-CoV, SARS-CoV or any types of viruses which has been discovered and contains their specific genome.
Introduction
Coronaviruses were discovered in the 1930s. [1] Arthur Schalk and M.C. Hawn figure out in 1931 a new respiratory infection (og Coronaviruses) in North Dakota. The infected chickens’ mortality rate was 40%–90%. [2] Fred Beaudette and Charles Hudson isolated and cultivated the infectious bronchitis virus which caused the disease.[3] In the 1940s, during the world war, two more animal coronaviruses, mouse hepatitis virus (MHV), and transmissible gastroenteritis virus (TGEV) were isolated.[4] It was not realized at the time that these three different viruses were related to coronavirus.[7] Impotently; human coronaviruses were discovered in the 1960s.[5-6] They were isolated using two different methods in the UK and the US.[8] E.C. Kendall, Malcom Byone, and David Tyrrell in 1960 isolated from a boy a new common cold virus B814.[9-11] The virus, unfortunately, was not able to be cultivated using standard techniques which had successfully cultivated rhinoviruses, adenoviruses, and other known common cold viruses. In 1965, Tyrrell and Byone [12] The new cultivating method was introduced to the lab by Bertil Hoorn. [13] The isolated virus when intranasally inoculated into volunteers caused a cold and was inactivated by ether which indicated it had a lipid envelope. [14-15] Around the same time, Dorothy Hamre [16] and John Procknow at the University of Chicago isolated 229E virus from medical students, which they grew in kidney tissue culture. The novel virus 229E, like the virus strain B814, when inoculated into volunteers caused a cold and was inactivated by ether. [17] B814 and 229E viruses were subsequently imaged by electron microscopy in 1967 by Scottish virologist June Almeida at St. Thomas Hospital in London. [18-19] Not only B814 and 229E viruses were they related to each other, but they were morphologically related to infectious bronchitis virus (IBV). [20] A research group at the National Institute of Health the same year was able to isolate another member of this new group of viruses using organ culture and named the virus strain OC43 (OC for organ culture).[21] Like B814, 229E, and IBV, the novel cold virus OC43 had distinctive club-like spikes when observed with the electron microscope.[22-23] The IBV-like novel cold viruses were shown to be morphologically related to the mouse hepatitis virus.[24] This new group of IBV-like viruses came to be known as coronaviruses after their distinctive morphological appearance.[25] Human coronavirus 229E and OC43 continued to be studied for decades. [26-27] The coronavirus strain B814 was lost. It is not known which present human coronavirus it was. [28] Other human coronaviruses discovered and named SARS-CoV in 2003, HCoV NL63 in 2004, HCoV HKU1 in 2005, MERS-CoV in 2012, and SARS-CoV-2 in 2019 (COVID-19). [29-30] There have also been a large number of animal coronaviruses identified since the 1960s which shows these strains of viruses are high in number and very difficult to be studied one by one. [31]
Origin
The most recent common ancestor (MRCA) of all coronaviruses is estimated to have existed as recently as 8000 BCE, although some models place the common ancestor as far back as 55 million years or more, implying long term coevolution with bat and avian species. [32] The most recent common ancestor of the alpha coronavirus line has been placed at about 2400 BCE, of the betacoronavirus line at 3300 BCE, of the gamma coronavirus line at 2800 BCE, and of the delta coronavirus line at about 3000 BCE. Bats and birds, as warm-blooded flying vertebrates, are an ideal natural reservoir for the coronavirus gene pool (with bats the reservoir for alpha coronaviruses and betacoronavirus – and birds the reservoir for gamma coronaviruses and delta coronaviruses). The large number and global range of bat and avian species that host viruses have enabled extensive evolution and dissemination of coronaviruses. [33]
Many human coronaviruses have their origin in bats.[34] The human coronavirus NL63 shared a common ancestor with a bat coronavirus (ARCoV.2) between 1190 and 1449 CE.[35] The human coronavirus 229E shared a common ancestor with a bat coronavirus (GhanaGrp1 Bt CoV) between 1686 and 1800 CE.[36] More recently, alpaca coronavirus and human coronavirus 229E diverged sometime before 1960.[68] MERS-CoV emerged in humans from bats through the intermediate host of camels.[37] MERS-CoV, although related to several bat coronavirus species, appears to have diverged from these several centuries ago.[38] The most closely related bat coronavirus and SARS-CoV diverged in 1986.[39] A possible path of evolution of SARS coronavirus and keen bat coronaviruses is that SARS-related coronaviruses coevolved in bats for a long time. The ancestors of SARS-CoV first infected leaf-nose bats of the genus Hipposideridae; subsequently, they spread to horseshoe bats in the species Rhinolophidae, then to civets, and finally to humans. [40-41]
Unlike other Betacoronaviruses, bovine coronavirus of the species Betacoronavirus 1 and subgenus Embecovirus is thought to have originated in rodents and not in bats.[42-43] In the 1790s, equine coronavirus diverged from the bovine coronavirus after a cross-species jump.[44] Later in the 1890s, human coronavirus OC43 diverged from bovine coronavirus after another cross-species spillover event.[45-46] It is speculated that the flu pandemic of 1890 may have been caused by this spillover event, and not by the influenza virus, because of the related timing, neurological symptoms, and unknown causative agent of the pandemic.[47] Besides causing respiratory infections, human coronavirus OC43 is also suspected of playing a role in neurological diseases.[48] In the 1950s, the human coronavirus OC43 began to diverge into its present genotypes.[49] Phylogenetically, mouse hepatitis virus (Murine coronavirus), which infects the mouse's liver and central nervous system, [50] is related to human coronavirus OC43 and bovine coronavirus. Human coronavirus HKU1, like the aforementioned viruses, also has its origins in rodents. [51] Well-known rodents include mice, rats, squirrels, prairie dogs, chipmunks, chinchillas, porcupines, beavers, guinea pigs, hamsters, gerbils, and capybaras. Rabbits, hares, and pikas.
Materials and Methods
SARS-CoV-2 is a complex virus. the genome of the virus differs from one person to another like fingerprints and the reason behind this phenomenon is the amount of tRNA and the metabolism (Mitochondrial Function) of the individual which is based on the amount of ATP releasing from the Mitochondria of each cell and loss of MMP (mitochondrial membrane permeabilization) which the reason behind the cause of cell apoptosis. Mitochondrial ATP generation requires proteins from the nuclear and mitochondrial genomes. ROS (Reactive Oxygen Species) disrupt the oxidative production of ATP, which is required for normal cellular function, due to the damage of mtDNA disrupts the normal synthesis of proteins needed for mitochondria function and making them suitable targets for attack by ROS produced during infections by viruses especially, any virus including Coronaviruses raises ROS in the host cell which affects Mitochondria and leads to loss of MMP. [52]
Proteins are translated by reading tri-nucleotides on the mRNA strand, also known as codons, from one end of the mRNA to the other (from the 5' to the 3' end). Each codon is translated into a single amino acid. Therefore, a shift of any number of nucleotides that is not divisible by 3 in the reading frame will result in subsequent codons to be read differently. [53] This effectively changes the ribosomal reading frame.
In viruses specifically COVID-19, Ribosomal Frameshift is programmed to allows the virus to encode multiple types of proteins from the same mRNA. HIV-1 (human immunodeficiency virus), [54] RSV (Rous sarcoma virus) [55], and all types of influenza viruses use Ribosomal Frameshift. they rely on frameshifting to create a proper ratio of normal translation and trans-frame (encoded by frameshifted sequence) proteins. Notably, its use in viruses is primarily for compacting more genetic information into a shorter amount of genetic material.
Effect of RNA Helicase in COVID-19
Helicases are enzymes that are vital to all living organisms. Their main function is to unpack an organism's genes. They are motor proteins that move directionally along a nucleic acid phosphodiester backbone, separating two annealed nucleic acid strands such as DNA and RNA using energy from ATP hydrolysis which proves that the vital COVID-19 is dependent on the amount of ATP production of the host cell by mitochondria. Approximately 1% of eukaryotic genes code for helicases. [56] The human genome codes for 95 non-redundant helicases: 64 RNA helicases (Which is important in infection of SARS-CoV-2) and 31 DNA helicases. Many cellular processes, such as DNA replication, transcription, translation, recombination, DNA repair, and ribosome biogenesis involve the separation of nucleic acid strands that necessitates the use of helicases.
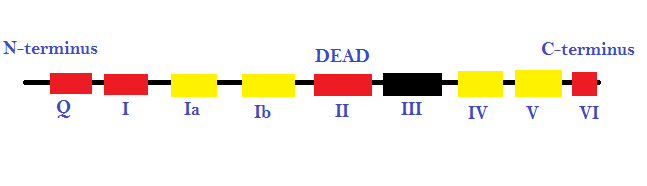 Figure 1: the different promoter sequences and accessory domains that aid in RNA unwinding (local strand separation). The regions in red are ATP binding domains and the regions in yellow are RNA interaction domains. Specific sequences termed DEAD-box proteins are also present that help catalyzes reactions in which ATP does not need to be directly hydrolyzed, as long as it binds to the domains on the strand.
Figure 1: the different promoter sequences and accessory domains that aid in RNA unwinding (local strand separation). The regions in red are ATP binding domains and the regions in yellow are RNA interaction domains. Specific sequences termed DEAD-box proteins are also present that help catalyzes reactions in which ATP does not need to be directly hydrolyzed, as long as it binds to the domains on the strand.
This image represents the different promoter sequences and accessory domains that aid in RNA unwinding (local strand separation). The regions in red are ATP binding domains and the regions in yellow are RNA interaction domains. Specific sequences termed DEAD-box proteins are also present that help catalyzes reactions in which ATP does not need to be directly hydrolyzed, as long as it binds to the domains on the strand.
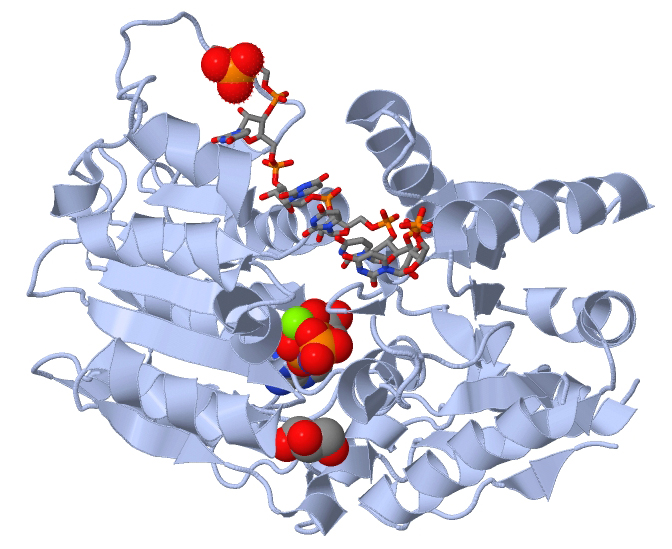
Figure 2: Human DEAD-box RNA helicase
Approximately 80% of all viruses are RNA viruses and they contain their specific RNA helicases. [58] Defective RNA helicases have been linked to infectious diseases (Viral Infections). [59] Some RNA helicases and DNA helicases can be found together in all the helicase super-families except for SF6. [60] [61] All the eukaryotic RNA helicases that have been identified up to date are non-ring forming and are part of SF1 and SF2. On the other hand, ring-forming RNA helicases have been found in bacteria and viruses. [62] RNA helicases that do exhibit unwinding activity have been characterized by at least two different mechanisms: canonical duplex unwinding and local strand separation. local strand separation occurs by a process wherein the helicase enzyme is loaded at any place along with the duplex. This is usually aided by a single-strand region of the RNA, and the loading of the enzyme is accompanied by ATP binding. [60] Once the helicase and ATP are bound, local strand separation occurs, which requires the binding of ATP but not the actual process of ATP hydrolysis. [62] Presented with fewer base pairs the duplex then dissociates without further assistance from the enzyme. This mode of unwinding is used by the DEAD/DEAH box helicases. [63]
|
RNA helicase |
|
|
Identifiers |
|
|
Databases |
|
|
PDB/structures |
|
Table 1: The table shows RNA Helicases [64] [65] [66]
In Vitro, to find the specific helicase activity of COVID-19, we used fluorescence-based assays, filtration assays, a scintillation proximity assay, a time-resolved fluorescence resonance energy transfer assay, and even used Trupoint diagnostic assay to observe the Helicase Activity. Even we have used the basic strand displacement assay which had been used in 1982–1983. [67] [68] The result was interestingly showed each Virus Helicase Assay is different from the other. The reason is the environment where the virus exits/evolves promptly and the only answer to this result is Lamarckian Evolution. As Coronaviruses (Including SarS-CoV-2) evolves very fast, their adaptation to the environment explains our results. [67] [68] [69]
Shifting balance theory, Bateson–Dobzhansky–Muller model & Quantum evolution
The Bateson–Dobzhansky–Muller model, [69] is a model of the evolution of genetic incompatibility, important in understanding the evolution of reproductive isolation during speciation and the role of natural selection in bringing it about. The theory was first described by William Bateson in 1909, [70] then by Theodosius Dobzhansky in 1934, and later by Herman Muller, H. Allen Orr, and Sergey Gavrilets. [71]. This model describes the drift between two species or even viruses to become hybrid and act differently in the environment. As many scientists only focus on the Co-Evolution of the viruses, Bateson–Dobzhansky–Muller model can be useful in describing the Fixation and Adaptation of SARS-CoV-2 in the different environment as well.
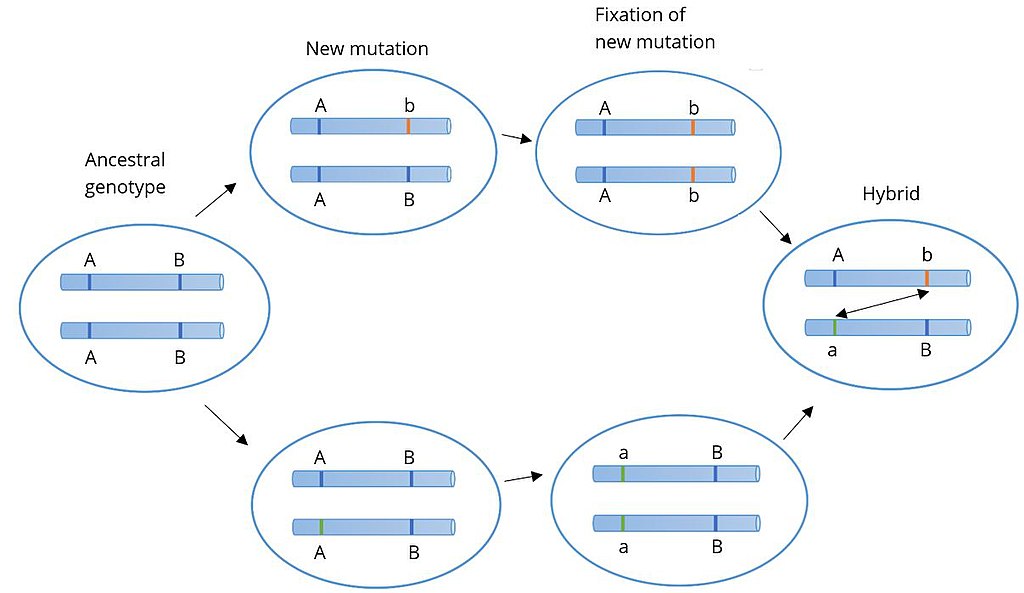
Figure 3: Bateson–Dobzhansky–Muller
each other, new mutations can arise. In one population A evolves into a, and in the other B evolves into b. When the two populations hybridize it is the first time a and b interact with each other. When these alleles are incompatible, we speak of Dobzhansky–Muller incompatibilities.
The Shifting balance theory is another theory of evolution proposed in 1932 by Sewall Wright, suggesting that adaptive evolution may proceed most quickly when a population of viruses divides into subpopulations with restricted gene flow. attempting to explain how a population may move across an adaptive valley to a higher adaptive peak. [72] According to the theory, this movement occurs in three steps:
All three steps describe adaption, Genetic Drift, and the Fitness of SARS-CoV-2. Novel Coronavirus has been evolved so fast from their ancestors to become a new Hybrid Novel RNA Virus and like cancer cells, they have rapid genetic mutations and adaptation to the environment.
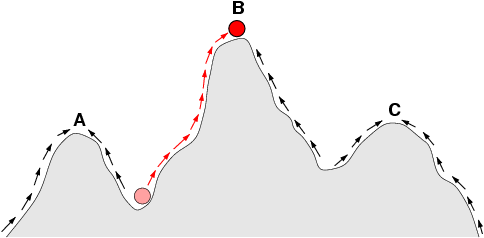
Figure 4: { The arrows indicate the preferred flow of a population (Viruses as well) on the landscape. The red ball indicates a population (or Viruses) that moves from an adaptive valley to the top of an adaptive peak. Based on natural selection (which usually acts to increase fitness in a population), it is not possible for a population at peak A to reach peak B because this requires descending into an adaptive valley. Shifting balance theory aims to explain how this may be possible.
Quantum evolution was proposed by George Gaylord Simpson in 1953. Quantum Evolution happens at the Taxonomic level and it plays a major role in the origin taxonomic units of relatively high rank in families, orders, and classes of species and parasites including viruses. As a whole, according to Simpson’s statements in 1944, quantum evolution resulted from Sewall Wright's model of random genetic drift. [74-83]
The History Behind the Neglected Coronavirus
We cannot forget the history of the outbreak of Coronaviruses. Firstly, it was considered harmless pathogens until they caused three major outbreaks of severe respiratory disease in the last 20 years. The Coronavirus was recognized in 1960 [84] and it was identified as a cause of the common cold. In 2002, it was considered as a nonfatal virus not severely pathogenic to humans. Then took place across the globe when an infected doctor traveled to Hong Kong in February 2003 [85] and transmitted the infection to other health workers and guests staying in the same hotel. These patients brought infection back to their home countries, that is, Singapore, Vietnam, and Canada. In the year 2012 [86], a new coronavirus emerged in the Middle East and was named Middle East Respiratory Syndrome (MERS). [86--90] Through our sessions above, the COVID-19 is a Hybrid Virus with a high mutation rate like cancer cells and has the potential of causing a new Hybrid Pandemic in the future. The Shifting Theory, Co-Evolution, Bateson–Dobzhansky–Muller model, and Quantum Evolution are the result of these high rare pandemics in the last 20 years.
Conclusion
Based on our research, to find the genome sequence of COVID-19 we also used Nanopore sequencing that introduced and manufactured by Oxford scientists, due to differences in the action of infection in the host, we could not reach any results since the Novel Virus has not a stable genome (which is quite dynamic) since through our deep research, each virus contains its specific genome sequencing and we cannot claim that COVID-19 has one specific genome sequence like MERS-CoV, SARS-C-V or any types of viruses which has been discovered and contains their specific genome. The main reason is the quick adaptation to the environment, Temperature, humidity, host genome type, host metabolism, Genetic Drift, Recombination of the virus, the high population of human beings, and the amount of ATP production of the host by their Mitochondria. Therefore; any type of vaccine cannot prevent the host from becoming infected by the virus. Vice versa, the new Coronavirus becomes more adapted and changes rapidly to make the environment and the host becoming weak and infected in the end.